|
 |
 |
 |
 |
 |
 |
 |
 |
 |

|
|
Recent invasion of DNA sequences into taxonomy is scary to some. While most researchers agree that sequence analysis is helpful, many feel somewhat lost because "feel and look" of DNA is different from that of specimens. In addition, probably because people enjoy arguing with each other, and because several classes of phylogenetic methods have been proposed (parsimony, distance, maximum likelihood, etc.), there is no agreement about how phylogenetic analysis of DNA should be done to be maximally productive. It seems obvious that all these classes of methods should be considered, as simplifying assumptions in each class and case may be quite different. However, due to various reasons, most research groups tend to pick their class of choice and criticize others who pick another class of methods. Typical, but not very useful.

Poladryas arachne arachne (W. H. Edwards, 1869), ♂
USA CO:Saguache Co., Noland Gulch, 27-V-09
© Kim Davis & Mike Stangeland
The idea behind this essay is to show that there is no magic in DNA analysis and it can make sense in easy ways. As a disclaimer, my apologies to those who understand evolutionary analysis of DNA is detail, as many concepts here are necessarily simplified, but all the simplifications are carefully considered to highlight the substance behind the form.
Genus Poladryas is an interesting North American genus of 2 species. Its phylogenetic affinities among the tribe Melitaeini were not well understood. We illustrate how analysis of DNA sequences can be used to clarify the position of Poladryas among its closest relatives. This serves as an example to describe the logic behind phylogenetic analysis of DNA.
Very impressive studies of the family Nympalidae have been performed recently by the Niklas Wahlberg group. The group maintains a wonderful web site: The Nymphalidae Systematics Group and pioneered application of major DNA techniques to Butterflies, including the polished protocol of data acquisition. Wahlberg and colleagues obtained partial DNA sequences for many species of Nymphalidae. Most of the published sequences are available from the GenBank database for free to everyone to analyze. Five of these sequences are used here.
For simplicity, we have chosen to analyze just one gene: 16S ribosomal RNA gene, which is a standard marker for phylogeny reconstruction in many groups of organisms. Other genes can be added to the analysis – you are welcome to try! However, the conclusions will not change.
The first step is to obtain needed sequences from the database. Submitting (Poladryas 16S ribosomal RNA gene) to the GenBank database search at http://www.ncbi.nlm.nih.gov/nuccore retrieves one entry: web page under this link. The entry contains a partial sequence of the gene. To get just the sequence only, go to FASTA link close to the top left of the NCBI web page. "FASTA" is a special sequence format, in which the first line is starting from the ">" character and is a description of the sequence. Sequence itself follows in other lines. You can cut'n'paste the retrieved sequence in your favorite text editor to work with it further. Here is how this FASTA-formatted sequence looks like:
>gi|8388955|gb|AF186854.1| Poladryas arachne voucher NW27-4 16S ribosomal RNA gene, partial sequence; mitochondrial TCAAAAACATGTCTTTTTGAAAATAATTTAAAGTTTAATCTGCCCACTGATATATTTATTAAAGGGCTGC AGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAAGACTTGTATGAAAGATTTGAT GAAATATAAACTGTCTCTAATTTAATAATAAAATTTAATTTTTTAGTTAAAAAGCTAAAATAATATTAAA AGACGAGAAGACCCTATAAAGTTTTATAATTTATTTATTTAATATTAAATATATAATTAATTATAGTAAT TATATAAAATTATTTTATTGGGGTGATAGAAAAATTTAATAAACTTTTTTTATATTATTAACATAAATAA GTGAAAAAATGATCCATTATTAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTTTT TTTAGAACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTAAATGCAAAAGTT TAAAATTTTGATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
We choose 4 other genera representing the 4 groups from the Melitaeini Tribe, namely Melitaea, Euphydryas, Chlosyne and Phyciodes. While the sequence for only one Poladryas species is available, all these 4 genera are represented in GenBank by many species. We have semi-randomly chosen the following well-known species: Melitaea athalia (common European species), Euphydryas phaeton, Chlosyne theona, and Phyciodes tharos. As it turned out, this choice worked well for the analysis. Here are GenBank sequences of all 5 taxa:
>gi|8388955|gb|AF186854.1| Poladryas arachne voucher NW27-4 16S ribosomal RNA gene, partial sequence; mitochondrial TCAAAAACATGTCTTTTTGAAAATAATTTAAAGTTTAATCTGCCCACTGATATATTTATTAAAGGGCTGC AGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAAGACTTGTATGAAAGATTTGAT GAAATATAAACTGTCTCTAATTTAATAATAAAATTTAATTTTTTAGTTAAAAAGCTAAAATAATATTAAA AGACGAGAAGACCCTATAAAGTTTTATAATTTATTTATTTAATATTAAATATATAATTAATTATAGTAAT TATATAAAATTATTTTATTGGGGTGATAGAAAAATTTAATAAACTTTTTTTATATTATTAACATAAATAA GTGAAAAAATGATCCATTATTAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTTTT TTTAGAACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTAAATGCAAAAGTT TAAAATTTTGATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT >gi|8388958|gb|AF186857.1| Melitaea athalia isolate 5-5 16S ribosomal RNA gene, partial sequence; mitochondrial TCAAAAACATGTCTTTTTGAAAATAATTTAAAGTCTAATCTGCCCACTGATATAATTATTAAAGGGCTGC AGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAAGACTTGTATGAAAGATTTGAT GAAATATATACTGTCTCTAATTTATGAATAAAAATTAATTTTTTAGTTAAAAAGCTAAAATAATATTAAA AGACGAGAAGACCCTATAAAGTTTTATAATTTATTTATTTAATATTAATTATATAAATAAATATAATAAT TAATTTAAATTATTTTATTGGGGTGATAAAAAAATTTAATTAACTTTTTTTAAAAAATAAACATAAATAA GTGTGATAATGATCCATTATTAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTTTT TTTAGTACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTAAGTGCAAAAGTT TAAAATTTTGATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT >gi|8389009|gb|AF186908.1| Euphydryas phaeton voucher NW13-3 16S ribosomal RNA gene, partial sequence; mitochondrial TCAAAAACATGCTTTTTATATTAATTTAAAGTCTAATCTGCCCACTGATAAATATTATTAAAGGGCTGCA GTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAAGACTTGTATGAAAGATTTGATG AAATATAAACTGTCTCTAATTTATAAATAGAAATTAATTTTTTAATTAAAAAGTTAAAATATTATTAAAA GACGAGAAGACCCTATAGAGTTTTATAATTTATTTATTTAATTATAAAATATATATTTAAATTTAATAAA TAAATAAATTATTATATTGGGGTGATAAAAAAATTTAATAAACTTTTTTTAATTAAATAACATAAATAAA TGAAAAAATGATCCATTATTAATGATTAGAAGAAAAAATTACCTTAGGGATAACAGCGTAATGTTTTTTT TTAGTACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTAAATGCAAAAGTTT AAAAGTTTTGATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT >gi|8389018|gb|AF186917.1| Chlosyne theona voucher NW27-6 16S ribosomal RNA gene, partial sequence; mitochondrial TCAAAAACATGTCTTTTTGATAATAATTTAAAGTCTAATCTGCCCACTGATATAATTATTAAAGGGCTGC AGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAAGACTTGAATGAAAGATTTGAT GAAATATAAACTGTCTCTAATTTAAAAATAAAATTTAATTTTTTAGTTAAAAAGCTAAAATATTATTAAA AGACGAGAAGACCCTATAAAGTTTTATAATTTATTTATTTAACATTAAATATATAATTAATTATGATAAT TAAATAAATTATTTTATTGGGGTGATAGAAAAATTTAATAAACTTTTTTTAAAAATAAACATAAATAAGT GAATAAATGATCCATTATTAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTTTTTT TAGTACAAATAAGAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTAAATGCCAAAATTTA AAATTTTAGATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT >gi|8389017|gb|AF186916.1| Phyciodes tharos voucher NW34-2 16S ribosomal RNA gene, partial sequence; mitochondrial TCAAAAACATGTCTTTTTGATAATAATTTAAAGTCTAATCTGCCCACTGATATAAATATTAAAGGGCTGC AGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAAGACTTGTATGAAAGATTTGAT GAAATATAATCTGTCTCTATATTATTAATAGAAATTAATTTTTTAGTTAAAAAGCTAAAATAGTATTAAA AGACGAGAAGACCCTATAAAGTTTTATAATTTATTTATTTATTATTAATTGTAAAAATAAATATTATAAT TAAATAAATTATTTTATTGGGGTGATAGAAAAATTAAATAAACTTTTTTTTAATATAAAACATAAATAAA TGAAAAATTGATCCATTAATAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTTTTT TTAGTACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTAAATGCAAAAGTTT AAAATTTTGATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
The second step is to align the sequences, i.e. to place letters (=nucleotides) in the same column if these letters are in evolutionarily equivalent positions of these sequences. Nucleotides in the same column = they all evolved from a common ancestral nucleotide. We use the MUSCLE server at EBI. Select "ClustalW2 (Strict)" for the "Output Format", instead of default "FASTA" and paste the above 5 sequences into the window. "Run" it and in less than a minute we get the result. Since sequences are long, the alignment is cut into blocks. Each block starts from the names of sequences taken from the first line (definition, starts with ">"). To obtain nice names (not numbers as in defines shown above), the defines were replaced with species names to get this alignment:
CLUSTAL W (1.81) multiple sequence alignment
Euphydryas_phaeton TCAAAAACATGCTTTTT--ATATTAATTTAAAGTCTAATCTGCCCACTGATAAATATTAT
Phyciodes_tharos TCAAAAACATGTCTTTTTGATAATAATTTAAAGTCTAATCTGCCCACTGATATA-AATAT
Melitaea_athalia TCAAAAACATGTCTTTTTGAAAATAATTTAAAGTCTAATCTGCCCACTGATATA-ATTAT
Chlosyne_theona TCAAAAACATGTCTTTTTGATAATAATTTAAAGTCTAATCTGCCCACTGATATA-ATTAT
Poladryas_arachne TCAAAAACATGTCTTTTTGAAAATAATTTAAAGTTTAATCTGCCCACTGATATA-TTTAT
*********** **** * * *********** ***************** * ***
Euphydryas_phaeton TAAAGGGCTGCAGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAA
Phyciodes_tharos TAAAGGGCTGCAGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAA
Melitaea_athalia TAAAGGGCTGCAGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAA
Chlosyne_theona TAAAGGGCTGCAGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAA
Poladryas_arachne TAAAGGGCTGCAGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAA
************************************************************
Euphydryas_phaeton GACTTGTATGAAAGATTTGATGAAATATAAACTGTCTCTAATTTATAAATAGAAATTAAT
Phyciodes_tharos GACTTGTATGAAAGATTTGATGAAATATAATCTGTCTCTATATTATTAATAGAAATTAAT
Melitaea_athalia GACTTGTATGAAAGATTTGATGAAATATATACTGTCTCTAATTTATGAATAAAAATTAAT
Chlosyne_theona GACTTGAATGAAAGATTTGATGAAATATAAACTGTCTCTAATTTAAAAATAAAATTTAAT
Poladryas_arachne GACTTGTATGAAAGATTTGATGAAATATAAACTGTCTCTAATTTAATAATAAAATTTAAT
****** ********************** ********* *** **** ** *****
Euphydryas_phaeton TTTTTAATTAAAAAGTTAAAATATTATTAAAAGACGAGAAGACCCTATAGAGTTTTATAA
Phyciodes_tharos TTTTTAGTTAAAAAGCTAAAATAGTATTAAAAGACGAGAAGACCCTATAAAGTTTTATAA
Melitaea_athalia TTTTTAGTTAAAAAGCTAAAATAATATTAAAAGACGAGAAGACCCTATAAAGTTTTATAA
Chlosyne_theona TTTTTAGTTAAAAAGCTAAAATATTATTAAAAGACGAGAAGACCCTATAAAGTTTTATAA
Poladryas_arachne TTTTTAGTTAAAAAGCTAAAATAATATTAAAAGACGAGAAGACCCTATAAAGTTTTATAA
****** ******** ******* ************************* **********
Euphydryas_phaeton TTTATTTATTTAATTATAAAATATATATTTAAATTTAATAAATAAAT-AAATTATTATAT
Phyciodes_tharos TTTATTTATTTAT-TATTAATTGTAAAAATAAATATTATAATTAAAT-AAATTATTTTAT
Melitaea_athalia TTTATTTATTTAA-TATTAATTATATAAATAAATATAATAATTAATTTAAATTATTTTAT
Chlosyne_theona TTTATTTATTTAA-CATTAAATATATAATTAATTATGATAATTAAAT-AAATTATTTTAT
Poladryas_arachne TTTATTTATTTAA-TATTAAATATATAATTAATTATAGTAATTATATAAAATTATTTTAT
************ ** ** * ** * *** * * *** ** * ******** ***
Euphydryas_phaeton TGGGGTGATAAAAAAATTTAATAAACTTTTTTTAATTAAATAACATAAATAAATGAAAAA
Phyciodes_tharos TGGGGTGATAGAAAAATTAAATAAACTTTTTTTTAATATAAAACATAAATAAATGAAAAA
Melitaea_athalia TGGGGTGATAAAAAAATTTAATTAACTTTTTTTAAAAAATAAACATAAATAAGTGTGATA
Chlosyne_theona TGGGGTGATAGAAAAATTTAATAAACTTTTTTTAAA-AATAAACATAAATAAGTGAATAA
Poladryas_arachne TGGGGTGATAGAAAAATTTAATAAACTTTTTTTATATTATTAACATAAATAAGTGAAAAA
********** ******* *** ********** *********** ** *
Euphydryas_phaeton ATGATCCATTATTAATGATTAGAAGAAAAAATTACCTTAGGGATAACAGCGTAATGTTTT
Phyciodes_tharos TTGATCCATTAATAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTT
Melitaea_athalia ATGATCCATTATTAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTT
Chlosyne_theona ATGATCCATTATTAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTT
Poladryas_arachne ATGATCCATTATTAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTT
********** ********* ************* ******************* ****
Euphydryas_phaeton TTTTTAGTACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTA
Phyciodes_tharos TTTTTAGTACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTA
Melitaea_athalia TTTTTAGTACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTA
Chlosyne_theona TTTTTAGTACAAATAAGAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTA
Poladryas_arachne TTTTTAGAACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTA
******* ******** *******************************************
Euphydryas_phaeton AATGCAAAAGTTTAAAAGTTTTGATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
Phyciodes_tharos AATGCAAAAGTTTAAAATTTT-GATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
Melitaea_athalia AGTGCAAAAGTTTAAAATTTT-GATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
Chlosyne_theona AATGCCAAAATTTAAAATTTTAGATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
Poladryas_arachne AATGCAAAAGTTTAAAATTTT-GATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
* *** *** ******* *** **************************************
We see that sequences are very similar, i.e. in many columns letters did not change. The last line in each alignment block (stars) marks positions that are identical in all 5 sequences. If there is no star, then at least one sequence contains a letter (=nucleotide) different from some other sequences at this position.
The third step is to reconstruct evolutionary tree from this alignment. As a first try we will use BioNJ program available as a web-server. Paste the above alignment – including the first line "CLUSTAL W (1.81) multiple sequence alignment" – in the window, put 1000 in the "Settings" field for "Number of bootstraps" and submit. Again, less than a minute is needed to get the result. The following tree in the "Newick format" will be generated:
This tree will look like this in ASCII form, if Euphydryas phaeton is taken as a root;
-----0.005----
+--------------------------------------------------------------------------------------------Euphydryas_phaeton
|
|
| +-------------------------------------------Chlosyne_theona
| +-----------------+
| | +---------------------------------------------------Poladryas_arachne
| +-------+
| | |
| | +-----------------------------------------------------------Melitaea_athalia
+-----------------+
|
+------------------------------------------------------------------------------------Phyciodes_tharos
Or like this in graphics form, shown by TreeView:
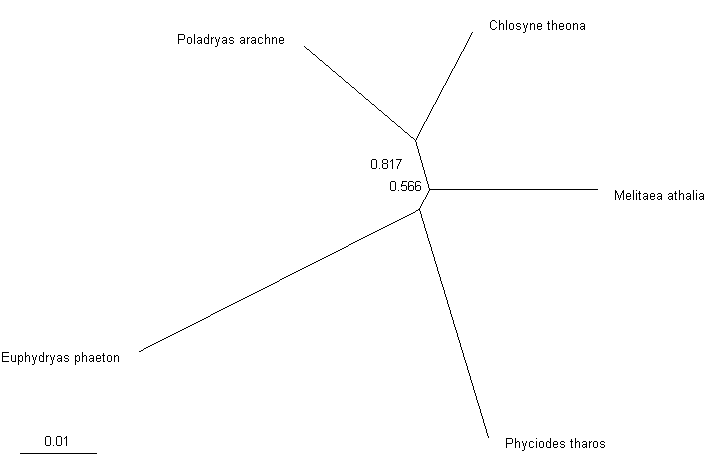
Unrooted tree of the 5 species build from fragments of 16S mitochondrial RNA sequences. Numbers by the nodes indicate
bootstrap support. The scale unit is the number of
expected nucleotide substitutions per site.
The same tree displayed by ATV and in a rooted form. The root position is chosen by the user.
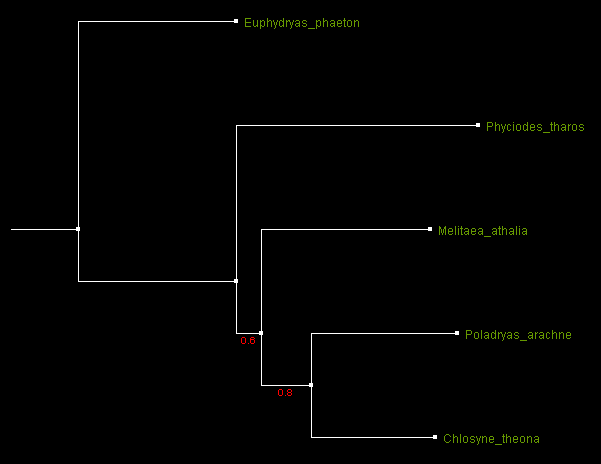
Rooted tree of the 5 species build from fragments of 16S mitochondrial RNA sequences. Numbers by the branches indicate
bootstrap support.
One can stop here and say that the analysis is done! Bootstrap support is a fractional value (between 0 and 1) that indicates how consistent positions in sequence are to support the tree. Values above 0.75 are considered reasonably strong, and in the tree above we see that Poladryas is grouped with Chlosyne. Mission accomplished! One can try to run other phylogenetic programs from the same web-server, e.g. PhyML, TNT, and MrBayes, or other programs, e.g. maximum parsimony dnapars, but the results will be the same. E.g. the TNT tree looks like this:
,-- Euphydryas_phaeton
|--| ,-- Phyciodes_tharos
`--| ,-- Melitaea_athalia
`--| ,-- Poladryas_arachne
`----- Chlosyne_theona
However, such a "black box" approach to obtain phylogeny does not help our understanding of how sequences are converted into a tree. Here we would like to explain how one can look at the letters in the alignment and analyze them manually.
Manual analysis. Let's look at the alignment again:
Euphydryas_phaeton TCAAAAACATGCTTTTT--ATATTAATTTAAAGTCTAATCTGCCCACTGATAAATATTAT
Phyciodes_tharos TCAAAAACATGTCTTTTTGATAATAATTTAAAGTCTAATCTGCCCACTGATATA-AATAT
Melitaea_athalia TCAAAAACATGTCTTTTTGAAAATAATTTAAAGTCTAATCTGCCCACTGATATA-ATTAT
Chlosyne_theona TCAAAAACATGTCTTTTTGATAATAATTTAAAGTCTAATCTGCCCACTGATATA-ATTAT
Poladryas_arachne TCAAAAACATGTCTTTTTGAAAATAATTTAAAGTTTAATCTGCCCACTGATATA-TTTAT
*********** **** * * *********** ***************** * ***
Euphydryas_phaeton TAAAGGGCTGCAGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAA
Phyciodes_tharos TAAAGGGCTGCAGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAA
Melitaea_athalia TAAAGGGCTGCAGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAA
Chlosyne_theona TAAAGGGCTGCAGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAA
Poladryas_arachne TAAAGGGCTGCAGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAA
************************************************************
Euphydryas_phaeton GACTTGTATGAAAGATTTGATGAAATATAAACTGTCTCTAATTTATAAATAGAAATTAAT
Phyciodes_tharos GACTTGTATGAAAGATTTGATGAAATATAATCTGTCTCTATATTATTAATAGAAATTAAT
Melitaea_athalia GACTTGTATGAAAGATTTGATGAAATATATACTGTCTCTAATTTATGAATAAAAATTAAT
Chlosyne_theona GACTTGAATGAAAGATTTGATGAAATATAAACTGTCTCTAATTTAAAAATAAAATTTAAT
Poladryas_arachne GACTTGTATGAAAGATTTGATGAAATATAAACTGTCTCTAATTTAATAATAAAATTTAAT
****** ********************** ********* *** **** ** *****
Euphydryas_phaeton TTTTTAATTAAAAAGTTAAAATATTATTAAAAGACGAGAAGACCCTATAGAGTTTTATAA
Phyciodes_tharos TTTTTAGTTAAAAAGCTAAAATAGTATTAAAAGACGAGAAGACCCTATAAAGTTTTATAA
Melitaea_athalia TTTTTAGTTAAAAAGCTAAAATAATATTAAAAGACGAGAAGACCCTATAAAGTTTTATAA
Chlosyne_theona TTTTTAGTTAAAAAGCTAAAATATTATTAAAAGACGAGAAGACCCTATAAAGTTTTATAA
Poladryas_arachne TTTTTAGTTAAAAAGCTAAAATAATATTAAAAGACGAGAAGACCCTATAAAGTTTTATAA
****** ******** ******* ************************* **********
Euphydryas_phaeton TTTATTTATTTAATTATAAAATATATATTTAAATTTAATAAATAAAT-AAATTATTATAT
Phyciodes_tharos TTTATTTATTTAT-TATTAATTGTAAAAATAAATATTATAATTAAAT-AAATTATTTTAT
Melitaea_athalia TTTATTTATTTAA-TATTAATTATATAAATAAATATAATAATTAATTTAAATTATTTTAT
Chlosyne_theona TTTATTTATTTAA-CATTAAATATATAATTAATTATGATAATTAAAT-AAATTATTTTAT
Poladryas_arachne TTTATTTATTTAA-TATTAAATATATAATTAATTATAGTAATTATATAAAATTATTTTAT
************ ** ** * ** * *** * * *** ** * ******** ***
Euphydryas_phaeton TGGGGTGATAAAAAAATTTAATAAACTTTTTTTAATTAAATAACATAAATAAATGAAAAA
Phyciodes_tharos TGGGGTGATAGAAAAATTAAATAAACTTTTTTTTAATATAAAACATAAATAAATGAAAAA
Melitaea_athalia TGGGGTGATAAAAAAATTTAATTAACTTTTTTTAAAAAATAAACATAAATAAGTGTGATA
Chlosyne_theona TGGGGTGATAGAAAAATTTAATAAACTTTTTTTAAA-AATAAACATAAATAAGTGAATAA
Poladryas_arachne TGGGGTGATAGAAAAATTTAATAAACTTTTTTTATATTATTAACATAAATAAGTGAAAAA
********** ******* *** ********** *********** ** *
Euphydryas_phaeton ATGATCCATTATTAATGATTAGAAGAAAAAATTACCTTAGGGATAACAGCGTAATGTTTT
Phyciodes_tharos TTGATCCATTAATAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTT
Melitaea_athalia ATGATCCATTATTAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTT
Chlosyne_theona ATGATCCATTATTAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTT
Poladryas_arachne ATGATCCATTATTAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTT
********** ********* ************* ******************* ****
Euphydryas_phaeton TTTTTAGTACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTA
Phyciodes_tharos TTTTTAGTACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTA
Melitaea_athalia TTTTTAGTACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTA
Chlosyne_theona TTTTTAGTACAAATAAGAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTA
Poladryas_arachne TTTTTAGAACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTA
******* ******** *******************************************
Euphydryas_phaeton AATGCAAAAGTTTAAAAGTTTTGATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
Phyciodes_tharos AATGCAAAAGTTTAAAATTTT-GATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
Melitaea_athalia AGTGCAAAAGTTTAAAATTTT-GATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
Chlosyne_theona AATGCCAAAATTTAAAATTTTAGATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
Poladryas_arachne AATGCAAAAGTTTAAAATTTT-GATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
* *** *** ******* *** **************************************
Positions marked with asterisks contain the same nucleotide in all 5 sequences. These positions are called invariant. Although these positions are quite important for various reasons, e.g. to compute nucleotide frequencies in the sequences, as these frequencies are need for some phylogenetic methods to work, invariant positions do not directly tell us which sequence is more similar to which other sequence. Thus, we can delete these positions from the alignment, and as a result, the alignment will be shorter and easier to view. Here is what we get:
Euphydryas_phaeton CT--TTCATATTAAATTAGAATTGATTAAATTTATAAAAA-AATAAATTAAATAAAAAATGCGTAAAGGTPhyciodes_tharos TCTGTACT-AATATTATTGAGCGAT-TTTGAAAAATATAA-TGAATAATATAAAAAAATAATATAAAGT-Melitaea_athalia TCTGAACT-ATTTAATTGAAGCAAA-TTTATAAAAAATATTTATTAAAAAATAGTGATATATATAGAGT-Chlosyne_theona TCTGTACT-ATAAAATAAATGCTAA-CTAATATTAGATAA-TGTAAAA-AATAGAATAATATATGACATAPoladryas_arachne TCTGAATT-TTTAAATATATGCAAA-TTAATATTAAGTTAATGTAATATTATTGAAAAATATAAAAAGT-The full alignment contained 540 positions, 470 out of which were invariant. Removal of them resulted in an alignment of 70 positions. Now, let's find positions that are semi-invariant, i.e. positions that are occupied by the same nucleotide in all but 1 sequence. Thus only one sequence will have a different nucleotide in these positions. This different nucleotide is marked gray in the alignment:
Euphydryas_phaeton CT--TTCATATTAAATTAGAATTGATTAAATTTATAAAAA-AATAAATTAAATAAAAAATGCGTAAAGGT 21Phyciodes_tharos TCTGTACT-AATATTATTGAGCGAT-TTTGAAAAATATAA-TGAATAATATAAAAAAATAATATAAAGT- 12Melitaea_athalia TCTGAACT-ATTTAATTGAAGCAAA-TTTATAAAAAATATTTATTAAAAAATAGTGATATATATAGAGT- 7Chlosyne_theona TCTGTACT-ATAAAATAAATGCTAA-CTAATATTAGATAA-TGTAAAA-AATAGAATAATATATGACATA 5Poladryas_arachne TCTGAATT-TTTAAATATATGCAAA-TTAATATTAAGTTAATGTAATATTATTGAAAAATATAAAAAGT- 7Number of nucleotides shaded gray is shown to the right of the alignment (highlighted yellow). We see that the number of gray marks is not the same in all sequences. In fact, Euphydryas phaeton sequence contains almost twice the number of such nucleotides than the next largest number (in Phyciodes tharos): 21 vs. 12. And 2-fold difference must mean something. Since all 5 sequences in the original alignment are about 90% identical, not that many nucleotide mutations happened in them. Therefore it is reasonable to think that each nucleotide difference usually corresponds to a single mutation in a gene. Since positions with gray highlight show difference in one species only, and all other species have the same nucleotide in these positions, it is reasonable to think that mutation to a different nucleotide happened on a path (=tree branch) leading to the species with the gray position. We can measure branch length in the number of mutations (=number of positions highlighted gray in each sequence). We see, that the branch leading to Euphydryas phaeton should be very long – almost twice the branch leading to Phyciodes tharos and at least three times longer than a branch leading to other species, i.e. at least about twice as many mutations happened on the branch leading to Euphydryas phaeton. Why is this branch so long?
Typically, if we consider very close organisms, most mutations separating them from each other are random and randomly happened in time, thus the more mutations, the more time separates the organisms. This idea is called "molecular clock". Of course, it is possible that during some times in evolution the "clock" might speed up, and in some species the rate of mutation accumulation might grow. However, why would Euphydryas phaeton accumulate mutations at a rate twice the rate of other species? Likely there is no reason. Thus a logical explanation for the length of this branch is that the root (common ancestor of the 5 genera) is placed on this branch. In other words, this branch consists of two branches: one is from the root (large blue circle) to Euphydryas phaeton and the other is from the root to the common ancestor of other 4 genera (small blue circle). With these 2 branches being about equal length (=about the same number of mutations accumulated during this time) we get an explanation why Euphydryas phaeton sequence shows about twice as many mutations as other sequences:
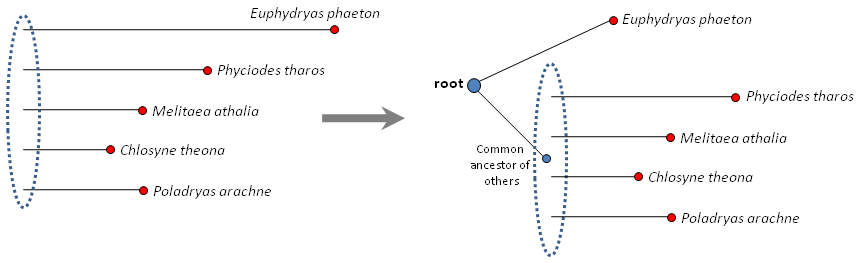
Starting from a completely unresolved tree with known terminal branch lengths (on the left) we deduce that the root is
likely to be on the longest branch.
Placing the root on the Euphydryas phaeton branch explains why this branch experienced many more mutations as other branches. Dashed ovals show yet unresolved
portions of the tree. Present day sequences are shown as red circles, the root is shown as a large blue circle, common ancestor of the 4 genera is shown as a small blue circle.
This root placement is the same, as the one proposed by The Nymphalidae Systematics Group on the basis of an outgroup – all other genera of Nymphalids and other butterflies. Now since we analyzed gray positions, let's remove them from the alignment and concentrate on the rest. We obtain 18 positions:
Euphydryas_phaeton TTAGATATAA-ATATAATPhyciodes_tharos TTTGAGTAAT-GTAAAA-Melitaea_athalia ATGAAATAAATAATAGA-Chlosyne_theona TAAATTATTG-G-TAGTAPoladryas_arachne AATATAATTAAGTTTGA-The next step is to check positions that have a unique nucleotide in a pair of sequences. Since we would like to check whether there is a sequence that is closest to the root, we mark positions that have the same nucleotide as the Euphydryas phaeton sequence. These positions are highlighted in red:
Euphydryas_phaeton TTAGATATAA-ATATAAT Phyciodes_tharos TTTGAGTAAT-GTAAAA- 3Melitaea_athalia ATGAAATAAATAATAGA- 1Chlosyne_theona TAAATTATTG-G-TAGTA 1Poladryas_arachne AATATAATTAAGTTTGA- 1We see that the last three sequences are consistently far from the Euphydryas phaeton sequence and share just 1 unique position with it, however, Phyciodes tharos has 3 such positions. Although 3 is not that different from 1, three ones in three other sequences are so consistent that we can form a hypothesis that Phyciodes tharos branched from the common ancestor next:
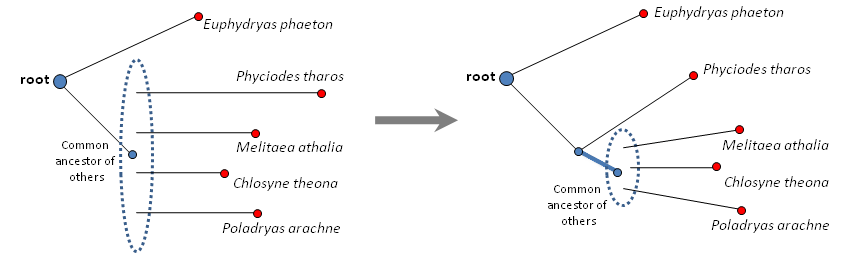
Resolving the tree further: blue thick branch signifies the 3 unique positions common to a pair Euphydryas phaeton and Phyciodes tharos:
these three mutations probably happened on this branch.
Since we are finished with the analysis of Euphydryas phaeton: it is quite clear that the root of the group falls on its branch, this sequence is removed from the alignment. Positions unique to each of the remaining sequences are highlighted in gray :
Phyciodes_tharos TTTGAGTAAT-GTAAAA- 3Melitaea_athalia ATGAAATAAATAATAGA- 1Chlosyne_theona TAAATTATTG-G-TAGTA 2Poladryas_arachne AATATAATTAAGTTTGA- 1Due to small number of unique positions, it is not certain how this analysis is helpful, however, the largest number of unique positions being in Phyciodes tharos is consistent with our hypothesis that this sequence branches out next in the tree. To concentrate on the rest of the alignment, we remove unique positions from it:
Phyciodes_tharos TTTAGTAAT-TMelitaea_athalia ATGAATAAATAChlosyne_theona TAATTATTG--Poladryas_arachne AATTAATTAATThen, analogously to the previous step, we highlight in red unique positions shared with Phyciodes tharos, which is the closest to the root:
Phyciodes_tharos TTTAGTAAT-T Melitaea_athalia ATGAATAAATA 5Chlosyne_theona TAATTATTG-- 2Poladryas_arachne AATTAATTAAT 2It is apparent that Melitaea athalia shares a larger number unique positions with Phyciodes tharos, and probably merged out next in the tree, leaving Chlosyne theona and Poladryas arachne as sister taxa. Additionally, we can look directly at Poladryas arachne sequence and check out unique positions it shares with other sequences, such positions are highlighted in green:
Phyciodes_tharos TTTAGTAAT-T 2 Melitaea_athalia ATGAATAAATA 3 Chlosyne_theona TAATTATTG-- 5 Poladryas_arachne AATTAATTAAT
The number of green positions is the largest in Chlosyne theona which is consistent with it being a sister taxon of Poladryas arachne. Thus we see that the following tree is reasonable. We can muddle it with statistics (and it will be fun!), but will refrain from it for now for the sake of the readers.
+----------- Euphydryas phaeton
|
root--| +-------- Phyciodes tharos
| |
+--+ +----- Melitaea athalia
| |
+--| +-- Chlosyne theona
+--+
+-- Poladryas arachne
Finally, it has to be noted that 3 positions supporting the branch between Euphydryas phaeton and Phyciodes tharos (blue thicker branch in a tree construction diagram above) is not a very large number of positions. It is conceivable that Phyciodes tharos and Melitaea athalia might be sister taxa. This, however, will not change the conclusion about Poladryas and Chlosyne being sister genera, as their common branch is supported by 5 positions.
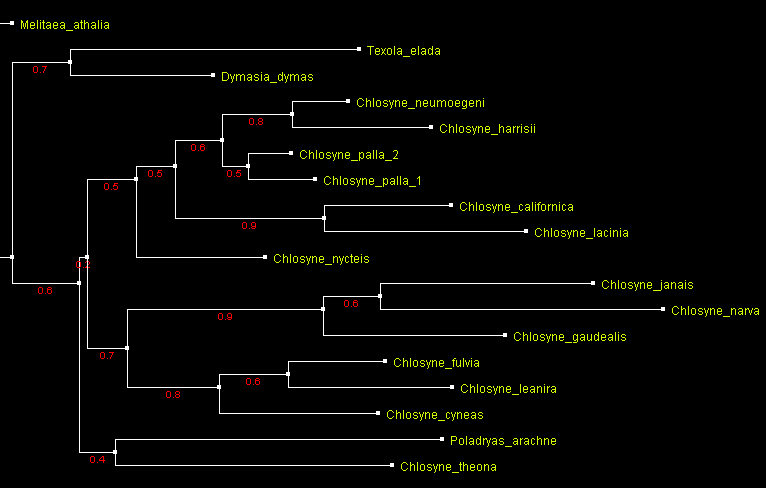
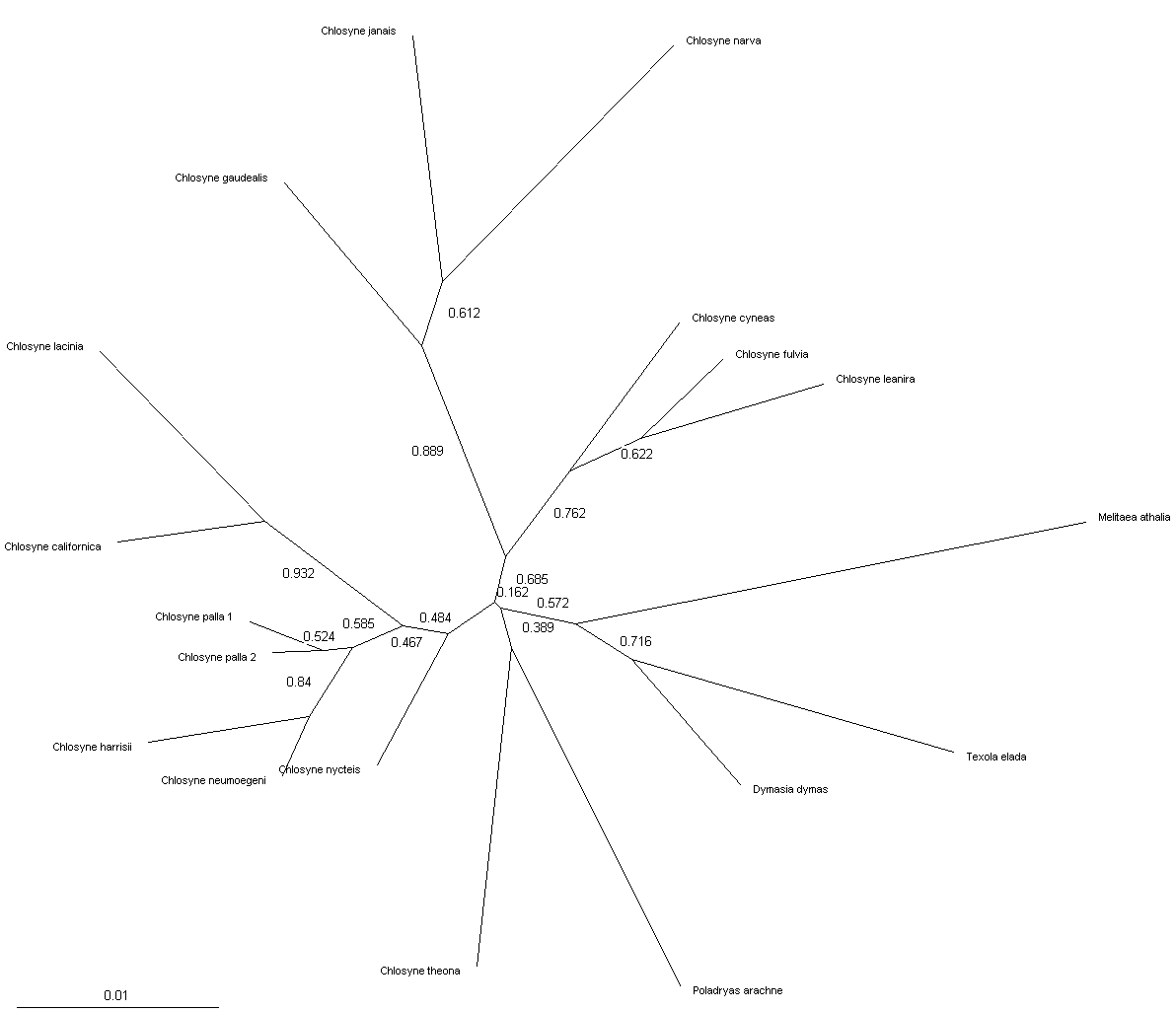
Just for fun – preliminary Chlosyne trees: