
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|
|
Phylogenetic reconstruction from DNA is a hard problem. Researchers complain that different genes and methods lead to inconsistent trees, morphological, biological and sequence data do not agree with each other very well, and many phylogenies remain unresolved despite serious effort. As a remedy, suggestion has been made to combine all the data available, e.g. concatenate several genes and add a sequence of morphological characters to this mix. This is a powerful remedy, because it makes sense that the use of all available data should be beneficial. However, questions remain about how to properly combine several genes (mutation rates in them can be quite different), or even worse, how to combine gene data with morphological data (e.g. how to weigh the characters in both sets relative to each other). Apparently, resulting phylogenies are sensitive to mutation rates and weights.
Here, we hypothesize that the major problem is not with the DNA sequences, incongruence of genes, or quality of phylogenetic methods. The problem is the lack of data. As it has been suggested several times before, if we had 10 to 100 fold more nucleotides known from each organisms, the major problem will disappear, and most observed incongruences will be a consequence of biology rather than the lack of data. While some researchers resort to simulations of DNA data obtained with pseudo-random number generator to support this hypothesis, we would like to work with real DNA sequences.
To illustrate this strongly expressed hypothesis we resort to the analysis of the Apes, namely genera Homo (human), Pan (chimp), Gorilla (gorilla), Pongo (orangutan) and Hylobates (gibbon). These primates receive much greater attention than insects, and more DNA sequences are available from them. Also, numerous studies have lead to the following consensus phylogeny, so we are quite confident in the correct tree:
+----------- Hylobates
|
root-----| +-------- Pongo
(e.g. | |
Macaca) +--+ +----- Gorilla
| |
+--| +-- Pan
+--+
+-- Homo
To simplify the problem even further, we focus on just one molecule – mitochondrial genome. Complete mitochondrial genomes are available for many Apes, and partial 16S ribosomal RNA sequences were obtained for many Nymphalidae species by Wahlberg and Zimmermann (2000). We can compare phylogeny reconstruction of the Apes using the segment corresponding to this 16S RNA fragment to the reconstruction from the complete mitochondrion DNA sequence. Why are the Apes a good model? Because genetic divergence as measured by the number of mutations per site is about the same between the 16S RNA fragments of the Apes and in some Nymphalidae tribes, such as Melitaeini. For instance the smallest distance of less than 1% is observed between the two Pan species, distance between Homo and Pan is about 4%, and the largest distance to Macaca mulatta (rhesus monkey) used as an outgroup is 13%. We expect that our conclusions might hold for the 16S ribosomal RNA sequences with divergence up to 15%.
Methods: All sequences used in this study were obtained from the GenBank database, and since mitochondrial genomes are circular, some sequence were circularly rearranged (permuted) to match others. Sequences were aligned using the MUSCLE server at EBI. The trees were reconstructed using the PhyML server with default parameters and visualized with TreeView and ATV. Tutorial about how to perform these procedure is available from here.
Data: DNA sequences for the Apes complete mitochondrial DNA, a fragment of 16S ribosomal RNA corresponding to the sequence know from Nymphalidae, multiple sequence alignment of complete mitochondrial DNA and of a fragment, trees based on complete and fragment sequences can be downloaded as text files.
Results: 16S RNA is a well-known phylogenetic marker, and its partial, 537 nucleotide sequences are available for Melitaeini (Nymphalidae), for instance for Poladryas arachne:
>gi|8388955|gb|AF186854.1| Poladryas arachne voucher NW27-4 16S ribosomal RNA gene, partial sequence; mitochondrial TCAAAAACATGTCTTTTTGAAAATAATTTAAAGTTTAATCTGCCCACTGATATATTTATTAAAGGGCTGC AGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATTGAAGACTTGTATGAAAGATTTGAT GAAATATAAACTGTCTCTAATTTAATAATAAAATTTAATTTTTTAGTTAAAAAGCTAAAATAATATTAAA AGACGAGAAGACCCTATAAAGTTTTATAATTTATTTATTTAATATTAAATATATAATTAATTATAGTAAT TATATAAAATTATTTTATTGGGGTGATAGAAAAATTTAATAAACTTTTTTTATATTATTAACATAAATAA GTGAAAAAATGATCCATTATTAATGATTAAAAGAAAAAATTACTTTAGGGATAACAGCGTAATATTTTTT TTTAGAACAAATAAAAAAAAAAGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTAAATGCAAAAGTT TAAAATTTTGATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
Is this ~500 nucleotide sequence sufficient for phylogeny reconstruction? Complete mitochondrial genome is known for the Apes and is 16,569 nucleotides long in human, which is about 30 times more nucleotides than the Nymphalidae sequences. Here is the human 16s RNA segment corresponding to the Poladryas arachne sequence, it is 572 nucleotides long:
>Homo_sapiens gi|251831106:2501-3072 Homo sapiens mitochondrion, complete genome CCAAAAACATCACCTCTAGCATCACCAGTATTAGAGGCACCGCCTGCCCAGTGACACATGTTTAACGGCC GCGGTACCCTAACCGTGCAAAGGTAGCATAATCACTTGTTCCTTAAATAGGGACCTGTATGAATGGCTCC ACGAGGGTTCAGCTGTCTCTTACTTTTAACCAGTGAAATTGACCTGCCCGTGAAGAGGCGGGCATAACAC AGCAAGACGAGAAGACCCTATGGAGCTTTAATTTATTAATGCAAACAGTACCTAACAAACCCACAGGTCC TAAACTACCAAACCTGCATTAAAAATTTCGGTTGGGGCGACCTCGGAGCAGAACCCAACCTCCGAGCAGT ACATGCTAAGACTTCACCAGTCAAAGCGAACTACTATACTCAATTGATCCAATAACTTGACCAACGGAAC AAGTTACCCTAGGGATAACAGCGCAATCCTATTCTAGAGTCCATATCAACAATAGGGTTTACGACCTCGA TGTTGGATCAGGACATCCCGATGGTGCAGCCGCTATTAAAGGTTCGTTTGTTCAACGATTAAAGTCCTAC GTGATCTGAGTT
Alignment of the two sequences reveals a reasonable degree of similarity, about 60% identity, so the fragments correspond to each other quite well:
CLUSTAL W (1.81) multiple sequence alignment
Poladryas_arachne TCAAAAACATGTCTTTTTG----AAAATAATTTAAAGTTTAATCTGCCCACTGATATATT
Homo_sapiens CCAAAAACATCACCTCTAGCATCACCAGTATTAGAGGCACCGCCTGCCCAGTGACACA--
********* * * * * * * *** * * ******* *** * *
Poladryas_arachne TATTAAAGGGCTGCAGTATATTGACTGTACAAAGGTAGCATAATCATTAGTCTTTTAATT
Homo_sapiens TGTTTAACGGCCGCGGTACCCTAACCGTGCAAAGGTAGCATAATCACTTGTTCCTTAAAT
* ** ** *** ** *** * ** ** ***************** * ** **** *
Poladryas_arachne GAAGACTTGTATGAAAGATTTGATGAAATATAAACTGTCTCTAA--TTTAATAATAAAAT
Homo_sapiens AGGGACCTGTATGAATGGCTCCACGAGGGTTCAGCTGTCTCTTACTTTTAACCAGTGAAA
*** ******** * * * ** * * ******** * ***** * **
Poladryas_arachne TTAATTTTTTAGTTAAAAAGCTAAAATAATATTAAAAGACGAGAAGACCCTATAAAGTTT
Homo_sapiens TTGACCTGCCCGTGAAGAGGCGGGCATAACACAGCAAGACGAGAAGACCCTATGGAGCTT
** * * ** ** * ** **** * ****************** ** **
Poladryas_arachne TATAATTTATTTATTTAA--------TATTAAATATATAATT----AATTATAGTAATTA
Homo_sapiens --TAATTTATTAATGCAAACAGTACCTAACAAACCCACAGGTCCTAAACTACCAAACCTG
********* ** ** ** *** * * * ** ** * *
Poladryas_arachne TATAAAATTATTTTATTGGGGTGAT-----AGAAAAATTTAATAAACTTTTTTTATATTA
Homo_sapiens CATTAAAAATTTCGGTTGGGGCGACCTCGGAGCAGAACCCAACCTCCGAGCAGTACATGC
** *** ** ****** ** ** * ** ** * ** **
Poladryas_arachne TTAACATAAATAAGTGAAA----------------AAATGATCCATTATTAATGATTAAA
Homo_sapiens TAAGACTTCACCAGTCAAAGCGAACTACTATACTCAATTGATCCAATAAC-TTGACCAAC
* * * * *** *** ** ******* ** *** **
Poladryas_arachne AGAAAAAATTACTTTAGGGATAACAGCGTAATATTTTTTTTTAGAACAAATAAAAAAAAA
Homo_sapiens GGAACAAGTTACCCTAGGGATAACAGCGCAATCCTATTCTAGAGTCCATATCAACAATAG
*** ** **** ************** *** * ** * ** ** ** ** ** *
Poladryas_arachne AGTTTGCGACCTCGATGTTGGATTAAGATAAAATTTAAATGCAAAAGTT-TAAAATTTTG
Homo_sapiens GGTTTACGACCTCGATGTTGGATCAGGACATCCCGATGGTGCAGCCGCTATTAAAGGTTC
**** ***************** * ** * **** * * * *** **
Poladryas_arachne ATCTGTTCGATCATTAAAATCTTACATGATCTGAGCT
Homo_sapiens GTTTGTTCAACGATTAAAGTCCTACGTGATCTGAGTT
* ***** * ****** ** *** ********* *
Sequences of equivalent fragments were extracted from the following species: Homo sapiens, Pan troglodytes and paniscus, Gorilla gorilla, Pongo abelii and pygmaeus, Hylobates lar and Macaca mulatta, which was used as an outgroup. Genetic divergence between these fragment sequences was as follows:
Macaca_mu Pongo_abe Pongo_pyg Hylobates Gorilla_g Homo_sapi Pan_trogl Pan_panis Macaca_mul 0.000000 0.119159 0.129449 0.129768 0.105026 0.115137 0.109002 0.107142 Pongo_abel 0.119159 0.000000 0.026632 0.098621 0.102064 0.084689 0.084589 0.084689 Pongo_pygm 0.129449 0.026632 0.000000 0.104649 0.100225 0.088654 0.094370 0.094481 Hylobates 0.129768 0.098621 0.104649 0.000000 0.088759 0.081129 0.084888 0.090807 Gorilla_go 0.105026 0.102064 0.100225 0.088759 0.000000 0.063365 0.054640 0.055903 Homo_sapie 0.115137 0.084689 0.088654 0.081129 0.063365 0.000000 0.042984 0.043033 Pan_troglo 0.109002 0.084589 0.094370 0.084888 0.054640 0.042984 0.000000 0.008793 Pan_panisc 0.107142 0.084689 0.094481 0.090807 0.055903 0.043033 0.008793 0.000000
Divergence covers the region up to 13% differences and seems appropriate as a model for butterfly taxa of Subfamily level, for instance divergence within Melitaeini falls well inside this interval as illustrated by this matrix of 8 taxa:
E.phaeton P.tharos C.lacinia P.arachne C.theona T.elada M.athalia M.didyma E. phaeton 0.000000 0.072449 0.076156 0.074205 0.069684 0.059372 0.069861 0.063456 P. tharos 0.072449 0.000000 0.067827 0.061716 0.053355 0.047121 0.051095 0.040996 C. lacinia 0.076156 0.067827 0.000000 0.050968 0.042814 0.046864 0.057319 0.053183 P. arachne 0.074205 0.061716 0.050968 0.000000 0.038811 0.049076 0.046879 0.044932 C. theona 0.069684 0.053355 0.042814 0.038811 0.000000 0.038840 0.044831 0.036832 T. elada 0.059372 0.047121 0.046864 0.049076 0.038840 0.000000 0.042836 0.032885 M. athalia 0.069861 0.051095 0.057319 0.046879 0.044831 0.042836 0.000000 0.026795 M. didyma 0.063456 0.040996 0.053183 0.044932 0.036832 0.032885 0.026795 0.000000
PhyML tree obtained from the alignment of 16S RNA fragments of 7 Ape species rooted with the rhesus monkey (Macaca mulatta) sequence is:
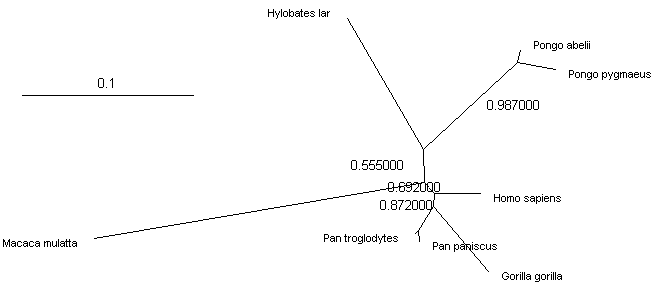
A tree of 7 Ape species built from fragments of 16S mitochondrial RNA sequences rooted with the rhesus monkey sequence
and displayed with TreeView.
Numbers by the nodes indicate
bootstrap support.
The scale unit is the number of
expected nucleotide substitutions per site.
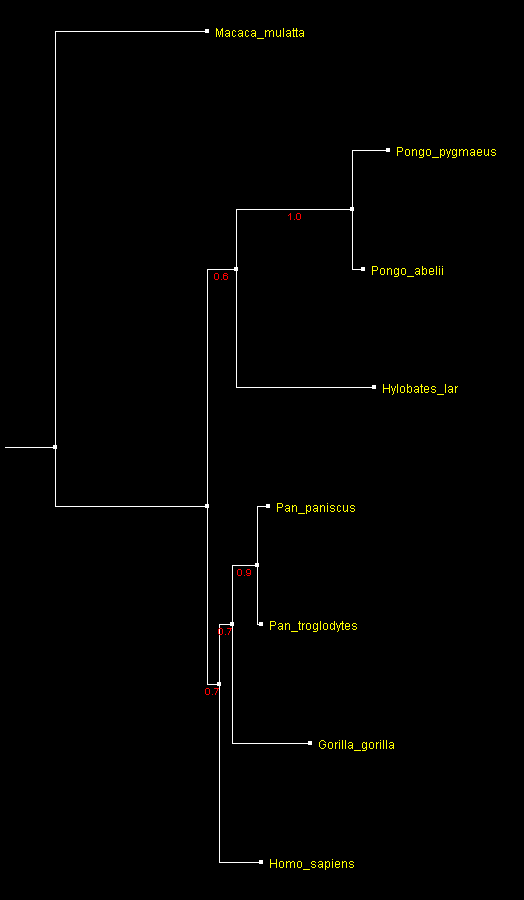
A tree of 7 Ape species built from fragments of 16S mitochondrial RNA sequences rooted with the rhesus monkey sequence
and displayed with ATV.
Numbers by the branches indicate
bootstrap support.
Apparently, this tree is not the same as the expected tree of the Apes shown at the top of this page. Pongo is grouped with Hylobates, and Pan is grouped with Gorilla, instead of forming a "ladder" tree. Obviously the two Pan and Pongo species are placed together correctly, and with strong support (values above 0.75 are more or less reliable). However, all other bootstrap values are below 0.75, and while the tree "looks good", it should be considered unresolved. Similar situation is observed for butterflies (Poladryas placement discussion). Since the Ape tree based on this fragment is incorrect, is it surprising that some of the butterfly trees do not appear consistent and sensible?
Next, we reconstruct the PhyML tree obtained from the alignment of complete mitochondrial genomes of the Apes. These genomes contain about 30 times more nucleotides than the fragment discussed above. Here is an example of a complete mitochondrial genome sequence:
>Pan_troglodytes gi|5835121|ref|NC_001643.1| Pan troglodytes mitochondrion, complete genome GTTTATGTAGCTTACCCCCTCAAAGCAATACACTGAAAATGTTTCGACGGGTTTACATCACCCCATAAAC AAACAGGTTTGGTCCTAGCCTTTCTATTAGCTCTTAGTAAGATTACACATGCAAGCATCCCCGCCCCGTG AGTCACCCTCTAAATCGCCATGATCAAAAGGAACAAGTATCAAGCACGCAGCAATGCAGCTCAAAACGCT TAGCCTAGCCACACCCCCACGGGAGACAGCAGTGATAAACCTTTAGCAATAAACGAAAGTTTAACTAAGC CATACTAACCTCAGGGTTGGTCAATTTCGTGCTAGCCACCGCGGTCATACGATTAACCCAAGTCAATAGA AACCGGCGTAAAGAGTGTTTTAGATCACCCCCCCATAAAGCTAAAATTCACCTGAGTTGTAAAAAACTCC AGCTGATACAAAATAAACTACGAAAGTGGCTTTAACACATCTGAATACACAATAGCTAAGACCCAAACTG GGATTAGATACCCCACTATGCTTAGCCCTAAACTTCAACAGTTAAATTAACAAAACTGCTCGCCAGAACA CTACGAGCCACAGCTTAAAACTCAAAGGACCTGGCGGTGCTTCATATCCCTCTAGAGGAGCCTGTTCTGT AATCGATAAACCCCGATCAACCTCACCGCCTCTTGCTCAGCCTATATACCGCCATCTTCAGCAAACCCTG ATGAAGGTTACAAAGTAAGCACAAGTACCCACGTAAAGACGTTAGGTCAAGGTGTAGCCTATGAGGTGGC AAGAAATGGGCTACATTTTCTACCCCAGAAAATTACGATAACCCTTATGAAACCTAAGGGTCAAAGGTGG ATTTAGCAGTAAACTAAGAGTAGAGTGCTTAGTTGAACAGGGCCCTGAAGCGCGTACACACCGCCCGTCA CCCTCCTCAAGTATACTTCAAAGGATACTTAACTTAAACCCCCTACGTATTTATATAGAGGAGATAAGTC GTAACATGGTAAGTGTACTGGAAAGTGCACTTGGACGAACCAGAGTGTAGCTTAACATAAAGCACCCAAC TTACACTTAGGAGATTTCAACTCAACTTGACCACTCTGAGCCAAACCTAGCCCCAAACCCCCTCCACCCT ACTACCAAACAACCTTAACCAAACCATTTACCCAAATAAAGTATAGGCGATAGAAATTGTAAACCGGCGC AATAGACATAGTACCGCAAGGGAAAGATGAAAAATTATACCCAAGCATAATACAGCAAGGACTAACCCCT GTACCTTTTGCATAATGAATTAACTAGAAATAACTTTGCAAAGAGAACCAAAGCTAAGACCCCCGAAACC AGACGAGCTACCTAAGAACAGCTAAAAGAGCACACCCGTCTATGTAGCAAAATAGTGGGAAGATTTATAG GTAGAGGCGACAAACCTACCGAGCCTGGTGATAGCTGGTTGTCCAAGATAGAATCTTAGTTCAACTTTAA ATTTACCTACAGAACCCTCTAAATCCCCTTGTAAACTTAACTGTTAGTCCAAAGAGGAACAGCTCTTTAG ACACTAGGAAAAAACCTTGTAAAGAGAGTAAAAAATTTAACACCCATAGTAGGCCTAAAAGCAGCCACCA ATTAAGAAAGCGTTCAAGCTCAACACCCACAACCTTAAAGATCCCAAACATACAACCGAACTCCTTACAC CCAATTGGACCAATCTATTACCCCATAGAAGAACTAATGTTAGTATAAGTAACATGAAAACATTCTCCTC CGCATAAGCCTACATCAGACCAAAATATTAAACTGACAATTAACAGCCTAATATCTACAATCAACCAACA AGCCATTATTACCCCCGCTGTTAACCCAACACAGGCATGCCCACAAGGAAAGGTTAAAAAAAGTAAAAGG AACTCGGCAAATCTTACCCCGCCTGTTTACCAAAAACATCACCTCTAGCATTACCAGTATTAGAGGCACC GCCTGCCCGGTGACATATGTTTAACGGCCGCGGTACCCTAACCGTGCAAAGGTAGCATAATCACTTGTTC CTTAAATAGGGACTTGTATGAATGGCTCCACGAGGGTTTAGCTGTCTCTTACTTTCAACCAGTGAAATTG ACCTACCCGTGAAGAGGCGGGCATAACATAACAAGACGAGAAGACCCTATGGAGCTTTAATTCATTAATG CAAACAATACTTAACAAACCTACAGGTCCTAAACTATTAAACCTGCATTAAAAATTTCGGTTGGGGCGAC CTCGGAGCACAACCCAACCTCCGAGCAATACATGCTAAGACCTCACCAGTCAAAGCGAATTACTACATCC AATTGATCCAATGACTTGACCAACGGAACAAGTTACCCTAGGGATAACAGCGCAATCCTATTCCAGAGTC CATATCAACAATAGGGTTTACGACCTCGATGTTGGATCAGGACATCCCGATGGTGCAGCCGCTATTAAAG GTTCGTTTGTTCAACGATTAAAGTCCTACGTGATCTGAGTTCAGACCGGAGTAATCCAGGTCGGTTTCTA TCTGTTCTAAATTTCTCCCTGTACGAAAGGACAAGAGAAATGAGGCCTACTTCACAAAGCGCCTTCCCCA ATAAATGATATTATCTCAATTTAGCGCCATGCCAACACCCACTCAAGAACAGAGTTTGTTAAGATGGCAG AGCCCGGTAATTGCATAAAACTTAAAACTTTACAATCAGAGGTTCAATTCCTCTTCTTGACAACACACCC ATGACCAACCTCCTACTCCTCATTGTACCCATCCTAATCGCAATAGCATTCCTAATGCTAACCGAACGAA AAATTCTAGGCTACATACAACTACGCAAAGGTCCCAACATTGTAGGTCCTTACGGGCTATTACAGCCCTT CGCTGACGCCATAAAACTCTTCACTAAAGAACCCTTAAAACCCTCCACTTCAACCATTACCCTCTACATC ACCGCCCCAACCCTAGCCCTCACCATTGCCCTCTTACTATGAACCCCCCTCCCCATACCCAACCCCCTAG TCAATCTTAACTTAGGCCTCCTATTTATTCTAGCCACCTCCAGCCTAGCCGTTTACTCAATCCTCTGATC AGGGTGAGCATCAAACTCGAACTACGCCTTAATCGGTGCACTACGAGCAGTAGCCCAAACAATCTCATAC GAAGTCACTCTAGCCATTATCCTACTGTCAACGCTACTAATAAGTGGCTCCTTCAATCTCTCTACCCTTG TCACAACACAAGAGCACCTCTGACTAATCCTGCCAACATGACCCCTGGCCATAATATGATTTATCTCTAC ACTAGCAGAGACCAACCGAACTCCCTTCGACCTTACTGAAGGAGAATCTGAACTAGTCTCAGGCTTTAAT ATCGAGTATGCCGCAGGCCCCTTTGCCCTATTTTTCATAGCCGAATACATAAACATTATTATAATAAACA CCCTCACTGCTACAATCTTCCTAGGAGCAACATACAATACTCACTCCCCTGAACTCTACACGACATATTT TGTCACCAAAGCTCTACTTCTAACCTCCCTGTTCCTATGAATTCGAACAGCATATCCCCGATTTCGCTAC GACCAGCTCATACACCTCCTATGAAAAAACTTCCTACCACTCACCCTAGCATCACTCATGTGATATATCT CCATACCCACTACAATCTCCAGCATCCCCCCTCAAACCTAAGAAATATGTCTGATAAAAGAATTACTTTG ATAGAGTAAATAATAGGAGTTCAAATCCCCTTATTTCTAGGACTATAAGAATCGAACTCATCCCTGAGAA TCCAAAATTCTCCGTGCCACCTATCACACCCCATCCTAAAGTAAGGTCAGCTAAATAAGCTATCGGGCCC ATACCCCGAAAATGTTGGTTACACCCTTCCCGTACTAATTAATCCCCTAGCCCAACCCATCATCTACTCT ACCATCCTTACAGGCACGCTCATTACAGCGCTAAGCTCACACTGATTTTTCACCTGAGTAGGCCTAGAAA TAAATATACTAGCTTTTATCCCAATCCTAACCAAAAAAATAAGCCCCCGCTCCACAGAAGCCGCCATCAA ATACTTTCTCACACAAGCAACTGCGTCCATAATTCTCCTGATAGCTATCCTCTCCAACAGCATACTCTCC GGACAATGAACCATAACCAATACTACCAATCAATACTCATCATTAATAATTATAATAGCAATGGCAATAA AACTAGGAATAGCCCCCTTTCACTTTTGAGTTCCAGAAGTTACCCAAGGCACCCCCCTAATATCCGGCCT ACTCCTCCTCACATGACAAAAATTAGCCCCTATTTCAATTATATACCAAATCTCCTCATCACTGAACGTA AACCTTCTCCTCACCCTTTCAATCTTGTCCATTATAGCAGGCAGCTGAGGCGGACTAAACCAAACCCAAC TACGCAAAATCCTAGCATACTCCTCAATCACCCACATAGGCTGAATAATAGCAGTCCTACCATATAACCC TAACATAACCATTCTTAATTTAACCATTTACATCATCCTAACTACTACCGCATTTCTGCTACTCAACTTA AACTCCAGCACCACAACCCTACTACTATCTCGCACCTGAAACAAGCTAACATGATTAACTCCCCTAATTC CATCCACCCTCCTCTCCCTAGGAGGCCTACCCCCACTAACTGGCTTCTTACCCAAATGAGTTATCATCGA AGAATTCACAAAAAATAATAGCCTCATCATCCCCACCATCATAGCCATCATCACTCTCCTTAACCTCTAT TTCTACCTACGCCTAATCTACTCCACCTCAATTACACTACTTCCCATATCTAATAACGTAAAAATAAAAT GACAATTCGAACATACAAAACCCACCCCCTTCCTCCCTACACTCATCACCCTTACCACACTGCTTCTACC CATCTCCCCCTTCATACTAATAATCTTATAGAAATTTAGGTTAAGCACAGACCAAGAGCCTTCAAAGCCC TCAGCAAGTTACAATACTTAATTTCTGCAACAACTAAGGACTGCAAAACCCCACTCTGCATCAACTGAAC GCAAATCAGCCACTTTAATTAAGCTAAGCCCTTACTAGATTAATGGGACTTAAACCCACAAACATTTAGT TAACAGCTAAACACCCTAATCAACTGGCTTCAATCTACTTCTCCCGCCGCAAGAAAAAAAGGCGGGAGAA GCCCCGGCAGGTTTGAAGCTGCTTCTTCGAATTTGCAATTCAATATGAAAATCACCTCAGAGCTGGTAAA AAGAGGCTTAACCCCTGTCTTTAGATTTACAGTCCAATGCTTCACTCAGCCATTTTACCCCACCCTACTG ATGTTCACCGACCGCTGACTATTCTCTACAAACCACAAAGATATTGGAACACTATACCTACTATTCGGTG CATGAGCTGGAGTCCTGGGCACAGCCCTAAGTCTCCTTATTCGGGCTGAACTAGGCCAACCAGGCAACCT CCTAGGTAATGACCACATCTACAATGTCATCGTCACAGCCCATGCATTCGTAATAATCTTCTTCATAGTA ATGCCTATTATAATCGGAGGCTTTGGCAACTGGCTAGTTCCCTTGATAATTGGTGCCCCCGACATGGCAT TCCCCCGCATAAACAACATAAGCTTCTGGCTCCTGCCCCCTTCTCTCCTACTTCTACTTGCATCTGCCAT AGTAGAAGCCGGCGCGGGAACAGGTTGAACAGTCTACCCTCCCTTAGCGGGAAACTACTCGCATCCTGGA GCCTCCGTAGACCTAACCATCTTCTCCTTACATCTGGCAGGCATCTCCTCTATCCTAGGAGCCATTAACT TCATCACAACAATTATTAATATAAAACCTCCTGCCATGACCCAATACCAAACACCCCTCTTCGTCTGATC CGTCCTAATCACAGCAGTCTTACTTCTCCTATCCCTCCCAGTCCTAGCTGCTGGCATCACCATACTATTG ACAGATCGTAACCTCAACACTACCTTCTTCGACCCAGCCGGGGGAGGAGACCCTATTCTATATCAACACT TATTCTGATTTTTTGGCCACCCCGAAGTTTATATTCTTATCCTACCAGGCTTCGGAATAATTTCCCACAT TGTAACTTATTACTCCGGAAAAAAAGAACCATTTGGATATATAGGCATGGTTTGAGCTATAATATCAATT GGCTTCCTAGGGTTTATCGTGTGAGCACACCATATATTTACAGTAGGGATAGACGTAGACACCCGAGCCT ATTTCACCTCCGCTACCATAATCATTGCTATTCCTACCGGCGTCAAAGTATTCAGCTGACTCGCTACACT TCACGGAAGCAATATGAAATGATCTGCCGCAGTACTCTGAGCCCTAGGGTTTATCTTTCTCTTCACCGTA GGTGGCCTAACCGGCATTGTACTAGCAAACTCATCATTAGACATCGTGCTACACGACACATACTACGTCG TAGCCCACTTCCACTACGTTCTATCAATAGGAGCTGTATTCGCCATCATAGGAGGCTTCATTCACTGATT CCCCCTATTCTCAGGCTATACCCTAGACCAAACCTATGCCAAAATCCAATTTGCCATCATGTTCATTGGC GTAAACCTAACCTTCTTCCCACAGCACTTCCTTGGCCTATCTGGGATGCCCCGACGTTACTCGGACTACC CCGATGCATACACCACATGAAATGTCCTATCATCCGTAGGCTCATTTATCTCCCTGACAGCAGTAATATT AATAATTTTCATGATTTGAGAAGCCTTTGCTTCAAAACGAAAAGTCCTAATAGTAGAAGAGCCCTCCGCA AACCTGGAATGACTATATGGATGCCCCCCACCCTACCACACATTCGAAGAACCCGTATACATAAAATCTA GACAAAAAAGGAAGGAATCGAACCCCCTAAAGCTGGTTTCAAGCCAACCCCATGACCTCCATGACTTTTT CAAAAAGATATTAGAAAAACTATTTCATAACTTTGTCAAAGTTAAATTACAGGTTAACCCCCGTATATCT TAATGGCACATGCAGCGCAAGTAGGTCTACAAGATGCTACTTCCCCTATCATAGAAGAACTTATTATCTT TCACGACCATGCCCTCATAATTATCTTTCTCATCTGCTTTCTAGTCCTATACGCCCTTTTCCTAACACTC ACAACAAAACTAACTAATACTAGTATTTCAGACGCCCAGGAAATAGAAACCGTCTGAACTATCCTGCCCG CCATCATCCTAGTCCTTATTGCCCTACCATCCCTGCGTATCCTTTACATAACAGACGAGGTCAACGACCC CTCCTTTACTATTAAATCAATCGGCCATCAATGATATTGAACCTACGAATACACCGACTACGGCGGGCTA ATCTTCAACTCCTACATACTCCCCCCATTATTTCTAGAACCAGGTGATCTACGACTCCTTGACGTTGATA ACCGAGTGGTCCTCCCAGTTGAAGCCCCCGTTCGTATAATAATTACATCACAAGATGTTCTACACTCATG AGCTGTTCCCACATTAGGCCTAAAAACAGACGCAATTCCCGGACGCCTAAACCAAACCACTTTCACCGCC ACACGACCAGGAGTATACTACGGCCAATGCTCAGAAATCTGTGGAGCAAACCACAGTTTTATACCCATCG TCCTAGAATTAATCCCTCTAAAAATCTTTGAAATAGGACCCGTATTCACTCTATAGCACCTTCTCTACCC CTCTCCAGAGCTCACTGTAAAGCTAACCTAGCATTAACCTTTTAAGTTAAAGATTAAGAGGACCGACACC TCTTTACAGTGAAATGCCCCAACTAAATACCGCCGTATGACCCACCATAATTACCCCCATACTCCTGACA CTATTTCTCGTCACCCAACTAAAAATATTAAATTCAAATTACCATCTACCCCCCTCACCAAAACCCATAA AAATAAAAAACTACAATAAACCCTGAGAACCAAAATGAACGAAAATCTATTCGCTTCATTCGCTGCCCCC ACAATCCTAGGCTTACCCGCCGCAGTACTAATCATTCTATTCCCCCCTCTACTGGTCCCCACTTCTAAAC ATCTCATCAACAACCGACTAATTACCACCCAACAATGACTAATTCAACTGACCTCAAAACAAATAATAAC TATACACAGCACTAAAGGACGAACCTGATCTCTCATACTAGTATCCTTAATCATTTTTATTACCACAACC AATCTTCTTGGGCTTCTACCCCACTCATTCACACCAACCACCCAACTATCTATAAACCTAGCCATGGCTA TCCCCCTATGAGCAGGCGCAGTAGTCATAGGCTTTCGCTTTAAGACTAAAAATGCCCTAGCCCACTTCTT ACCGCAAGGCACACCTACACCCCTTATCCCCATACTAGTTATCATCGAAACTATTAGCCTACTCATTCAA CCAATAGCCTTAGCCGTACGTCTAACCGCTAACATTACTGCAGGCCACCTACTCATGCACCTAATTGGAA GCGCCACACTAGCATTATCAACTATCAATCTACCCTATGCACTCATTATCTTCACAATTCTAATCCTACT GACTATTCTAGAGATCGCCGTCGCCTTAATCCAAGCCTACGTTTTTACACTTCTAGTGAGCCTCTACCTG CACGACAACACATAATGACCCACCAATCACATGCCTACCACATAGTAAAACCCAGCCCATGACCCCTAAC AGGGGCCCTCTCGGCCCTCCTAATAACCTCCGGCCTGGCCATATGATTCCACTTCTACTCCACAACACTA CTCACACTAGGCTTACTAACTAACACATTGACCATATATCAATGATGACGCGATGTTATACGAGAAGGCA CATACCAAGGCCACCACACACCACCCGTCCAAAAAGGTCTCCGATATGGGATAATTCTTTTTATTACCTC AGAAGTTTTTTTCTTTGCAGGATTTTTTTGAGCTTTCTACCACTCCAGCCTAGCCCCTACCCCCCAGCTA GGAGGACACTGGCCCCCAACAGGTATTACCCCACTAAATCCCCTAGAAGTCCCACTCCTAAACACATCTG TATTACTCGCATCAGGAGTATCAATTACTTGAGCCCATCACAGCTTAATAGAAAATAACCGAAACCAAAT AATTCAAGCACTGCTTATTACGATTCTACTAGGTCTTTATTTTACCCTCCTACAAGCCTCAGAATATTTC GAATCCCCTTTTACCATTTCCGATGGCATCTACGGCTCAACATTCTTTGTAGCCACAGGCTTCCACGGAC TCCACGTCATTATTGGATCAACTTTCCTCACTATCTGCCTCATCCGCCAACTAATATTTCACTTCACATC CAAACATCACTTCGGCTTTCAAGCCGCCGCCTGATACTGACACTTCGTAGATGTAGTCTGACTATTTCTA TATGTCTCTATTTACTGATGAGGATCTTACTCTTTTAGTATAAGTAGTACCGTTAACTTCCAATTAACTA GTTTTGACAACATTCAAAAAAGAGTAATAAACTTCGTCCTAATTTTAATAACCAATACCCTTCTAGCCCT ACTACTGATAATTATCACATTCTGACTACCACAACTCAACAGCTACATAGAAAAATCTACCCCTTACGAA TGTGGCTTCGACCCTATATCCCCCGCCCGCGTCCCCTTCTCCATAAAATTTTTCCTAGTAGCCATCACCT TCCTATTATTTGACCTAGAAATTGCCCTCCTATTGCCCTTACCTTGAGCCCTACAAACGGCCAACCTACC ACTAATAGTCACATCATCCCTCTTATTAATTACTATCCTAGCCCTAAGCCTCGCCTACGAATGATTACAA AAAGGGTTAGACTGAACCGAATTGGTATATAGTTTAAATAAAACGAATGATTTCGACTCATTAAATTATG ATAATCATATTTACCAAATGCCCCTTATTTATATAAATATTATACTAGCATTTACCATCTCACTTCTAGG AATACTAGTATATCGCTCACACCTAATATCTTCCCTACTATGCCTAGAAGGAATAATACTATCACTGTTC ATCATAGCCACCCTCATAACCCTCAATACTCACTCCCTCTTAGCCAATATTGTACCCATCACCATACTAG TCTTTGCTGCCTGCGAAGCAGCAGTAGGTCTAGCACTACTAGTTTCAATCTCTAACACATATGGCTTAGA CTACGTACATAACCTAAACCTACTCCAATGCTAAAACTAATCATCCCGACAATTATATTACTACCACTAA CATGATTCTCTAAAAAACGTATAATTTGAATCAACACAACCACTCACAGCCTAATTATCAGCACCATTCC CTTACTATTTTTTAACCAAATTAACAACAACCTATTCAGCTGTTCCCTGCCCTTCTCCTCCGACCCCTTA ACAACTCCCCTCCTAATATTAACTGCTTGACTTCTACCCCTCACAATCATAGCAAGCCAGCGCCACCTAT CCAACGAACCACTATCACGAAAAAAACTCTACCTCTCCATGCTAATTTCCCTCCAAATCTCCTTAATTAT AACATTCTCGGCCACAGAGCTAATTATATTTTATATCTTCTTCGAAACCACACTTATCCCCACCCTGGCT ATCATCACCCGATGGGGTAACCAACCAGAACGCCTGAACGCAGGTACATACTTCCTATTCTATACCCTAG TAGGCTCCCTCCCCCTACTCATCGCACTAATCTATACCCACAACACCCTAGGCTCACTAAATATCCTATT ACTCACTCTTACAACCCAAGAACTATCAAACACCTGAGCCAACAACTTAATATGACTAGCGTACACGATG GCTTTCATGGTAAAAATACCCCTTTACGGACTCCACCTATGACTCCCTAAAGCCCATGTCGAAGCCCCTA TTGCCGGGTCAATGGTACTTGCTGCAGTACTCTTAAAATTAGGTGGCTATGGCATAATACGCCTCACACT CATCCTCAACCCCCTAACAAAACATATAGCCTATCCCTTCCTCATGTTGTCCTTATGAGGTATAATCATA ACAAGCTCCATCTGCCTGCGACAAACAGACCTAAAATCGCTCATTGCATACCCTTCAGTCAGCCACATAG CCCTCGTAGTAACAGCCATTCTCATCCAAACCCCCTGAAGCTTCACCGGCGCAATTATCCTCATAATCGC CCACGGACTTACATCCTCATTATTATCCTGCCTAGCAAACTCAAATTATGAACGCACCCACAGTCGCATC ATAATTCTCTCCCAAGGACTTCAAACTCTACTCCCACTAATAGCCTTTTGATGACTCCTGGCAAGCCTCG CTAACCTCGCCCTACCCCCTACCATTAATCTCCTAGGGGAACTCTCCGTGCTAGTAACCTCATTCTCCTG ATCAAATACCACTCTCCTACTCACAGGATTCAACATACTAATCACAGCCCTGTACTCCCTCTACATGTTT ACCACAACACAATGAGGCTCACTCACCCACCACATTAATAGCATAAAGCCCTCATTCACACGAGAAAACA CTCTCATATTTTTACACCTATCCCCCATCCTCCTTCTATCCCTCAATCCTGATATCATCACTGGATTCAC CTCCTGTAAATATAGTTTAACCAAAACATCAGATTGTGAATCTGACAACAGAGGCTCACGACCCCTTATT TACCGAGAAAGCTTATAAGAACTGCTAACTCGTATTCCCATGCCTAACAACATGGCTTTCTCAACTTTTA AAGGATAACAGTTATCCATTGGTCTTAGGCCCCAAAAATTTTGGTGCAACTCCAAATAAAAGTAATAACC ATGTATGCTACCATAACCACCTTAGCCCTAACTTCCTTAATTCCCCCCATCCTCGGCGCCCTCATTAACC CTAACAAAAAAAACTCATACCCCCATTACGTGAAATCCATTATCGCATCCACCTTTATCATTAGCCTTTT CCCCACAACAATATTCATATGCCTAGACCAAGAAACTATTATCTCGAACTGACACTGAGCAACAACCCAA ACAACCCAACTCTCCCTGAGCTTTAAACTAGACTATTTCTCCATAACATTTATCCCCGTAGCACTGTTCG TTACATGATCCATCATAGAATTCTCACTATGATATATAGACTCAGACCCCAACATCAACCAATTCTTCAA ATACTTACTTATCTTCCTAATTACTATACTAATCCTAGTCACCGCTAACAACCTATTCCAACTCTTCATC GGCTGAGAAGGCGTAGGAATTATATCCTTTCTACTCATTAGCTGATGGTACGCCCGAACAGATGCCAACA CAGCAGCCATCCAAGCAATCCTATATAACCGTATCGGTGATATTGGTTTTGTCCTAGCCCTAGCATGATT TCTCCTACACTCCAACTCATGAGATCCACAACAAATAATCCTCCTAAGTACTAATACAGACCTTACTCCA CTACTAGGCTTCCTCCTAGCAGCAGCAGGCAAATCAGCTCAACTAGGCCTTCACCCCTGACTCCCCTCAG CCATAGAAGGCCCTACCCCTGTTTCAGCCCTACTCCACTCAAGCACCATAGTCGTAGCAGGAATCTTCCT ACTCATCCGCTTCTACCCCCTAGCAGAGAATAACCCACTAATCCAAACTCTCACGCTATGCCTAGGCGCT ATCACCACCCTATTCGCAGCAGTCTGCGCCCTCACACAAAATGACATCAAAAAAATCGTGGCCTTCTCCA CTTCAAGCCAACTAGGACTCATAATAGTTACAATCGGTATCAACCAACCACACCTAGCATTCCTTCACAT CTGCACCCACGCTTTCTTCAAAGCCATACTATTCATATGCTCCGGATCCATTATTCACAACCTCAATAAT GAGCAAGACATTCGAAAAATAGGAGGATTACTCAAAACCATACCCCTCACTTCAACCTCCCTCACCATTG GGAGCCTAGCATTAGCAGGAATACCCTTCCTCACAGGTTTCTACTCCAAAGACCTCATCATCGAAACCGC TAACATATCATACACAAACGCCTGAGCCCTATCTATTACTCTCATCGCCACCTCTCTGACAAGCGCCTAC AGCACCCGAATAATCCTCCTCACCCTAACAGGTCAACCTCGCTTCCCAACCCTCACCAACATTAACGAAA ACAACCCCACTCTGTTAAATCCCATTAAACGCCTAACCATTGGAAGCTTATTTGCAGGATTTCTCATTAC CAACAACATTCTCCCCATATCTACTCCCCAAGTGACAATTCCCCTTTACTTAAAACTTACAGCCCTAGGC GTTACTTCCCTAGGACTTCTAACAGCCCTAGACCTCAATTACCTAACCAGCAAGCTCAAAATAAAATCCC CACTATATACATTTCACTTCTCTAATATACTCGGATTCTACCCTAACATTATACACCGCTCGATCCCCTA TCTAGGCCTTCTTACAAGCCAAAACCTACCCCTACTTCTTCTAGACCTGACCTGACTAGAGAAACTATTA CCTAAAACAATTTCACAGTACCAAATCTCCGCTTCCATTACCACCTCAACCCAAAAAGGCATGATCAAAC TTTATTTCCTCTCTTTTTTCTTCCCTCTCATCTTAACCTTACTCCTAATCACATAACCTATTCCCCCGAG CAATCTCAATCACAATGTATACACCAACAAACAATGTCCAACCAGTAACTACTACTAACCAACGCCCATA ATCATATAAGGCCCCCGCACCAATAGGATCCTCCCGAATCAGCCCTGGCCCCTCCCCTTCATAAATTATT CAACTTCCCACGCTATTAAAATTTACCACAACCACCATCCCATCATACCCTTTTACCCATAACACTAATC CTACCTCCATCGCCAGTCCTACTAAAACACTAACCAAAACCTCAACCCCTGACCCCCATGCCTCAGGATA CTCCTCAATAGCCATAGCCGTAGTATACCCAAAAACAACCATTATTCCCCCCAAATAAATTAAAAAAACC ATTAAACCTATATAACCTCCCCCATAATTCAAAATGATGGCACACCCAACTACACCACTAACAATCAATA CTAAACCCCCATAAATGGGAGAAGGCTTAGAAGAAAACCCCACAAACCCTATCACTAAACTCACACTCAA TAAAAATAAAGCATATGTCATTATTCTCGCACGGACTACAACCACGACCAATGATATGAAAAACCATCGT TGTATTTCAACTACAAGAACACCAATGACCCCGACACGCAAAATTAACCCACTAATAAAATTAATTAATC ACTCATTTATCGACCTCCCCACCCCATCCAACATTTCCGCATGATGGAACTTCGGCTCACTTCTCGGCGC CTGCCTAATCCTTCAAATTACCACAGGATTATTCCTAGCTATACACTACTCACCAGACGCCTCAACCGCC TTCTCGTCGATCGCCCACATCACCCGAGACGTAAACTATGGTTGGATCATCCGCTACCTCCACGCTAACG GCGCCTCAATATTTTTTATCTGCCTCTTCCTACACATCGGCCGAGGTCTATATTACGGCTCATTTCTCTA CCTAGAAACCTGAAACATTGGCATTATCCTCTTGCTCACAACCATAGCAACAGCCTTTATGGGCTATGTC CTCCCATGAGGCCAAATATCCTTCTGAGGAGCCACAGTAATTACAAACCTACTGTCCGCTATCCCATACA TCGGAACAGACCTGGTCCAGTGAGTCTGAGGAGGCTACTCAGTAGACAGCCCTACCCTTACACGATTCTT CACCTTCCACTTTATCTTACCCTTCATCATCACAGCCCTAACAACACTTCATCTCCTATTCTTACACGAA ACAGGATCAAATAACCCCCTAGGAATCACCTCCCACTCCGACAAAATTACCTTCCACCCCTACTACACAA TCAAAGATATCCTTGGCTTATTCCTTTTCCTCCTTATCCTAATGACATTAACACTATTCTCACCAGGCCT CCTAGGCGATCCAGACAACTATACCCTAGCTAACCCCCTAAACACCCCACCCCACATTAAACCCGAGTGA TACTTTCTATTTGCCTACACAATCCTCCGATCCATCCCCAACAAACTAGGAGGCGTCCTCGCCCTACTAC TATCTATCCTAATCCTAACAGCAATCCCTGTCCTCCACACATCCAAACAACAAAGCATAATATTTCGCCC ACTAAGCCAACTGCTTTACTGACTCCTAGCCACAGACCTCCTCATCCTAACCTGAATCGGAGGACAACCA GTAAGCTACCCCTTCATCACCATCGGACAAATAGCATCCGTATTATACTTCACAACAATCCTAATCCTAA TACCAATCGCCTCTCTAATCGAAAACAAAATACTTGAATGAACCTGCCCTTGTAGTATAAACTAATACAC CGGTCTTGTAAACCGGAAACGAAAACTTTCTTCCAAGGACAAATCAGAGAAAAAGTAATTAACTTCACCA TCAGCACCCAAAGCTAAGATTCTAATTTAAACTATTCTCTGTTCTTTCATGGGGAAGCAAATTTAGGTAC CACCTAAGTACTGGCTCATTCATTACAACCGCTATGTATTTCGTACATTACTGCCAGCCACCATGAATAT CGTACAGTACCATATCACCCAACTACCTATAGTACATAAAATCCACTCCCACATCAAAACCTTCACTCCA TGCTTACAAGCACGCACAACAATCAACTCCCAACTGTCGAACATAAAACACAATTCCAACGACACCCCTC CCCCACCCCGATACCAACAGACCTATCTCCCCTTGACAGAACATAGTACATACAACCATACACCGTACAT AGCACATTACAGTCAAACCCCTCCTCGCCCCCACGGATGCTCCCCCTCAGATAGGAATCCCTTGGTCACC ATCCTCCGTGAAATCAATATCCCGCACAAGAGTGACTCTCCTCGCTCCGGGCCCATAACATCTGGGGGTA GCTAAAGTGAACTGTATCCGACATCTGGTTCCTACCTCAGGGCCATGAAGTTCAAAAGACTCCCACACGT TCCCCTTAAATAAGACATCACGATGGATCACAGGTCTATCACCCTATTAACCAGTCACGGGAGCCTTCCA TGCATTTGGTATTTTCGTCTGGGGGGTGTGCACGCGATAGCATTGCGAAACGCTGGCCCCGGAGCACCCT ATGTCGCAGTATCTGTCTTTGATTCCTGCCCCATTGTATTATTTATCGCACCTACGTTCAATATTACGAC CTAGCATACCTACTAAAGTGTGTTGATTAATTAATGCTTGCAGGACATAACAACAGCAGCAAAATGCTCA CATAACTGCTTTCCACACCAACATCATAACAAAAAATTCCCACAAACCCCCCCTTCCCCCCGGCCACAGC ACTCAAACAAATCTCTGCCAAACCCCAAAAACAAAGAACCCAGACGCCAGCCTAGCCAGACTTCAAATTT CATCTTTAGGCGGTATGCACTTTTAACAGTCACCCCTCAATTAACATGCCCTCCCCCCTCAACTCCCATT CTACTAGCCCCAGCAACGTAACCCCCTACTCACCCTACTCAACACATATACCGCTGCTAACCCCATACCC TGAACCAACCAAACCCCAAAGACACCCCTACACA
The result is the following tree:

A tree of 7 Ape species built from complete mitochondrial DNA sequences rooted with the rhesus monkey sequence
and displayed with TreeView.
Numbers by the nodes indicate
bootstrap support.
The scale unit is the number of
expected nucleotide substitutions per site.
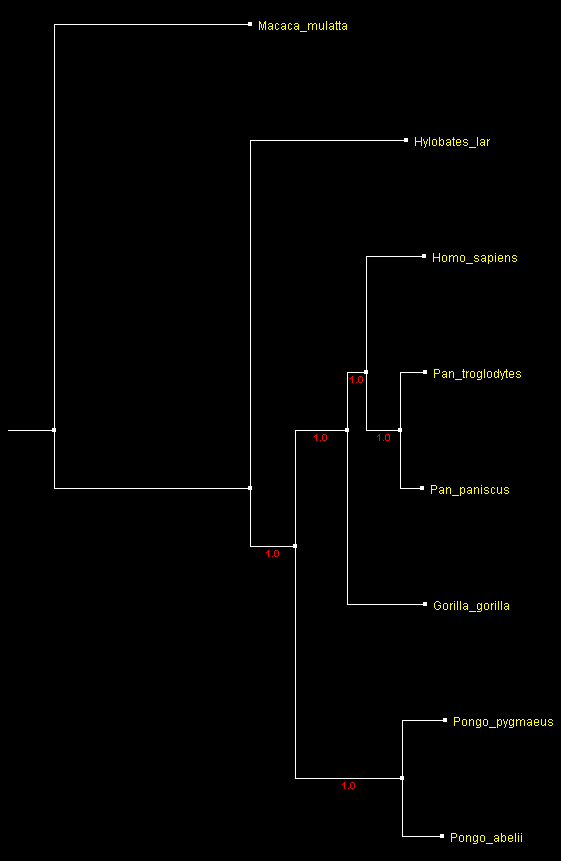
A tree of 7 Ape species built from complete mitochondrial DNA sequences rooted with the rhesus monkey sequence
and displayed with ATV.
Numbers by the branches indicate
bootstrap support.
Now, all the bootstrap values are at a 100% and the tree corresponds exactly to the expected tree of primates, in which Homo and Pan are the sister genera and others form a "ladder" to the root. 30 times more information leads to an excellent resolution of phylogeny from a single, although very long, DNA molecule without the use of any morphological characters and sophisticated tree reconstruction software.
While it is not surprising that short DNA sequences are insufficient for correct and confident phylogeny reconstruction, and, as we illustrate, longer sequences can lead to fully resolved tree, the question that cannot be answered here is whether such complete resolution will be obtained in all, or even in most cases. We suspect, however, that complete mitochondrial genomes will be a very powerful tool of phylogenetic reconstruction and will clarify many problems that are currently experienced by this relative young field of DNA-based taxonomy.
To show a counter-example, in which 16,500 nucleotides of mitochondrial genome is not enough to obtain confidently supported completely bipartitioned tree, we have chosen a recent study of rhinoceroses (Willerslev et al. 2009). The PhyML tree obtained from the alignment of complete mitochondrial sequences of rhinoceroses in agreement with the work of Willerslev et al. (2009) does not reveal the branching order and places White (Ceratotherium simum) and Black (Diceros bicornis) rhinos as the basal clade with a very weak support (22.3%). All other bipartitions are supported at a 100%. This interesting trichotomy is probably the result of divergence between the three rhino clades within a very short time period. One other interesting fact about this work is that woolly rhinoceros (Coelodonta antiquitatis) is currently extinct, and its mitochondrial genome was sequenced from a hair shaft of a fossil excavated from the permafrost in Yakutia (Russia).

A tree of 6 rhinoceros species built from complete mitochondrial DNA sequences rooted with the horse and tapir sequences
and displayed with TreeView.
Numbers by the nodes indicate
bootstrap support. The scale unit is the number of
expected nucleotide substitutions per site.

A tree of 6 rhinoceros species built from complete mitochondrial DNA sequences rooted with the horse and tapir sequences
and displayed with ATV.
Numbers by the branches indicate
bootstrap support.
The short tree branch separating the three clades of rhinos that cannot be resolved was estimated to span about 1 million years, and the age of the last common ancestor of all 6 rhinos is about 30 million years. Apparently, 16,000 nucleotides are not enough to provide resolution for 1 Myr time interval. Can researchers obtain data to resolve speciation events separated by 1 Myr? The suggestion has been made to collect nuclear sequences, or even complete genomes, to address this question.
Although 1 Myr branch at a time-distance of 30 Myr from today is not possible to resolve using mitochondrial genome, a more recent events separated by 1 Myr at a time-distance of about 7 Myr usually can be resolved if appropriate outgroup is available. The following example is particularly amazing, as mitochondrial genome of one prehistoric animal was needed to resolve the tree position of another prehistoric animal.
Woolly mammoth (Mammuthus primigenius) sequences were obtained recently and offered a puzzle whether mammoth is a sister species of Asian (Elephas maximus) or African (Loxodonta africana) elephants. The problem is due to short time-span during which Elephantidae species diverged (about 1 Myr) and the absence of close outgroup. Some of the closest present-day species are dugong (Dugong dugon) and hyrax (Procavia capensis). Using mitochondrial genomes of all 5 species we obtain alignment and PhyML tree that reveals no strong support (bootstrap ~0.4, and values below 0.75 are not indicative) and weakly groups mammoth with African elephant:

A tree of 2 present day elephant species and a woolly mammoth built from complete mitochondrial DNA sequences rooted with the dugong and hyrax sequences
and displayed with TreeView.
Numbers by the nodes indicate
bootstrap support. The scale unit is the number of
expected nucleotide substitutions per site.

A tree of 2 present day elephant species and a woolly mammoth built from complete mitochondrial DNA sequences rooted with the dugong and hyrax sequences
and displayed with ATV.
Numbers by the branches indicate
bootstrap support.
Apparently, closer outgroup or more sequences are needed to resolve mammoth position. However, no extant animals are closer to elephants than dugong and hyrax. Only extinct Elephantidae may offer a solution. Rohland et al. (2007) obtained complete mitochondrial genome sequence of American mastodon (Mammut americanum) from a tooth found in Alaska. Mastodons diverged from other Proboscideans about 25 Myr and the ratio of the number of transitions to the number of transversions between them has not reached saturation. Adding the mastodon sequence to the alignment results in the following tree:
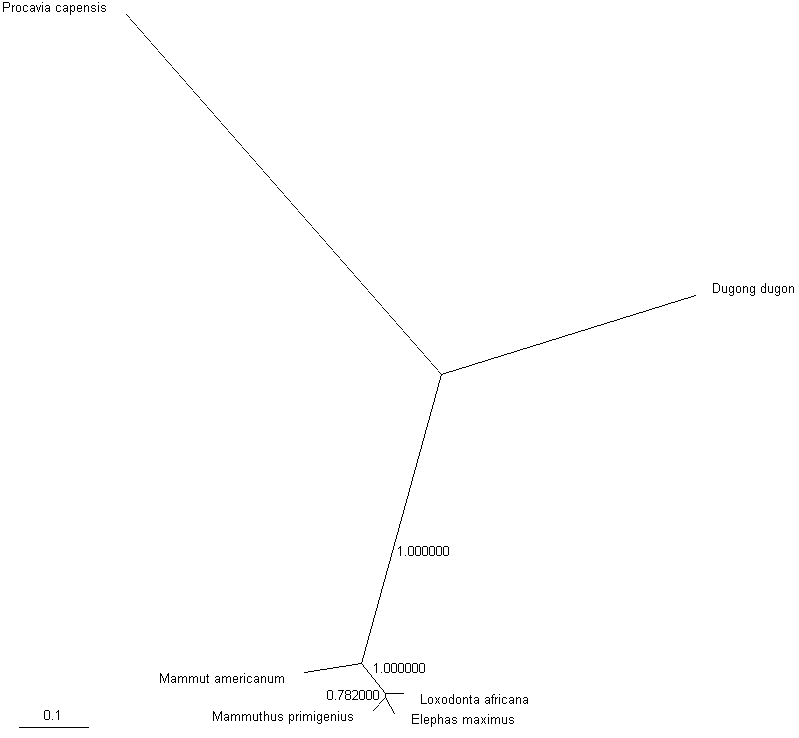
A tree of 2 present day elephant species, a woolly mammoth and an American mastodon built from complete mitochondrial DNA sequences rooted with the dugong and hyrax sequences
and displayed with TreeView.
Numbers by the nodes indicate
bootstrap support. The scale unit is the number of
expected nucleotide substitutions per site.
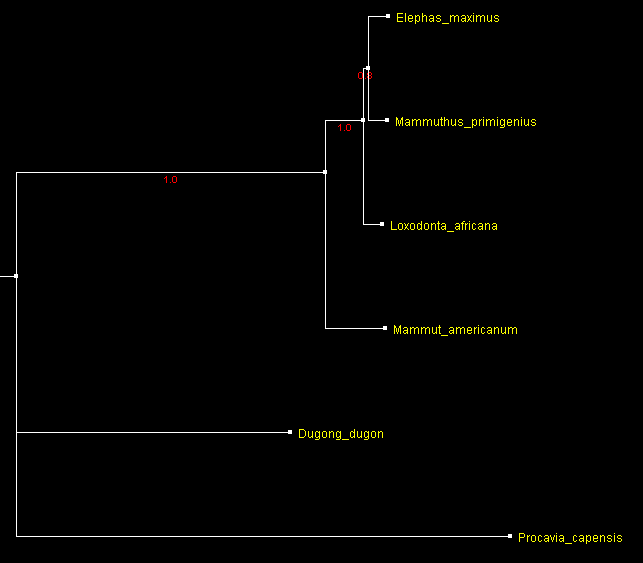
A tree of 2 present day elephant species, a woolly mammoth and an American mastodon built from complete mitochondrial DNA sequences rooted with the dugong and hyrax sequences
and displayed with ATV.
Numbers by the branches indicate
bootstrap support.
This trees shows moderate support (bootstrap about 0.8) for grouping mammoth with Asian elephant. Apparently, closer outgroup sequence (mastodon) changed the grouping in the tree. However, the presence of distant sequences from dugong and hyrax might be detrimental. Their removal shows the tree (build from the alignment of 4 sequence) with a high support (>0.9) for this grouping.

A tree of 2 present day elephant species and a woolly mammoth built from complete mitochondrial DNA sequences rooted with the mastodon sequence
and displayed with TreeView.
Number by the branch indicates
bootstrap support. The scale unit is the number of
expected nucleotide substitutions per site.
Can the {Asian elephant – mammoth} clade be supported even stronger without obtaining more sequences of other extinct animal species? Adding complete mitochondrion genome sequences of more specimens (2 specimens each) from these species provides close to 100% support for this short branch representing about 1 Myr even with dugong and hyrax sequences being present. The tree build from the alignment of these 9 sequence is the ultimate result:
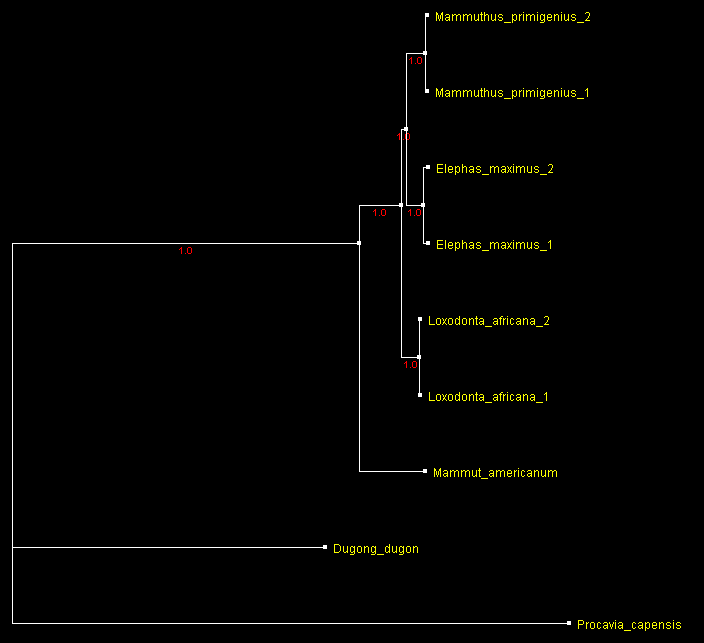
A tree of 2 present day elephant species and woolly mammoth (2 specimens of each) built from complete mitochondrial DNA sequences rooted with an American mastodon, dugong and hyrax sequences
and displayed with ATV.
Numbers by the branches indicate
bootstrap support.
Checking the tree build from the alignment of 8 sequence without the mastodon sequence with the hopes that sequences of specimen pairs will help resolution does not produce desired result: the tree is different, and the support for the branch of interest is very small (~0.3).

A tree of 2 present day elephant species and woolly mammoth (2 specimens of each) built from complete mitochondrial DNA sequences rooted with dugong and hyrax sequences
and displayed with ATV.
Numbers by the branches indicate
bootstrap support.
Conclusions: